The Annual Meeting of the American Association for Cancer Research (AACR) took place from April 17–22 in San Diego. This year’s theme, “Precision, Partnership, Purpose: Advancing Cancer Science to Save Lives Globally,” reflected the extraordinary advances in cancer research, as well as their expected impacts for patients across the globe. Several NICR research groups also presented their findings.
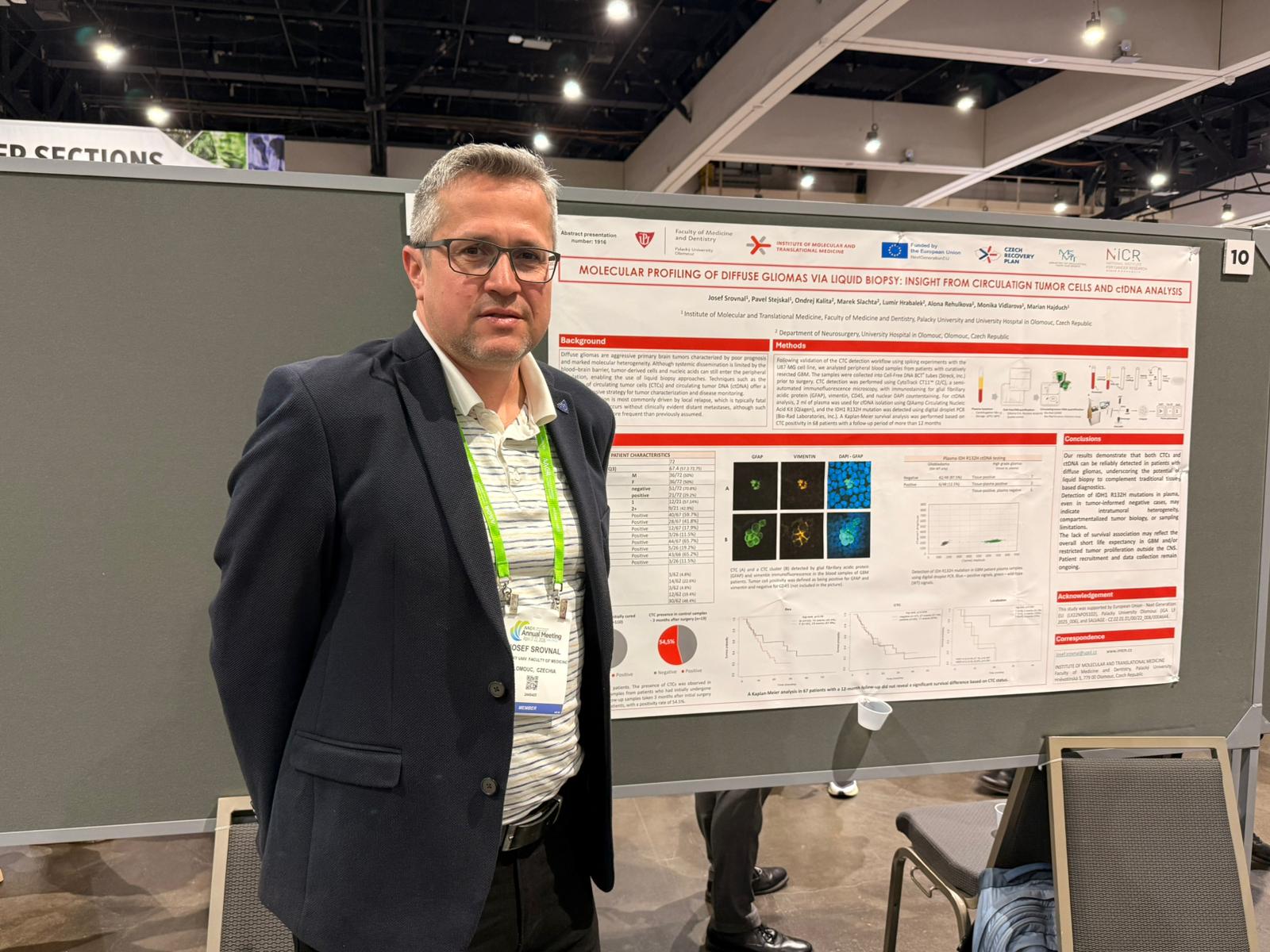
Molecular profiling of diffuse gliomas via liquid biopsy
Scientists from the research groups from the Olomouc node of the NICR evaluated the feasibility of molecular profiling of diffuse gliomas through the analysis of circulating tumour cells (CTCs) and circulating tumour DNA (ctDNA) in peripheral blood samples collected prior to surgery. ctDNA was analysed for the IDH1 R132H. CTCs were detected in 29.1% of samples obtained at first surgery and in 54.5% of samples from patients with recurrent disease. In a subgroup of 67 patients with IDH-wildtype glioblastoma, CTC status was not associated with overall survival. Plasma analysis identified IDH1 R132H mutations in 14% of patients, including some tumour-informed negative cases. These findings show that both CTCs and ctDNA can be detected in diffuse gliomas and may complement tissue-based molecular diagnostics. Detection of plasma IDH1 R132H despite tumour-informed negativity may reflect intratumoural heterogeneity, biological compartmentalisation, or sampling limitations. Recruitment and data collection are ongoing. (Abstract 1070. DOI: 10.1158/1538-7445.AM2026-1070)
Autophagy-modulating compounds and their off-target activities
Autophagy helps many tumours survive stress, metabolic pressure, and therapy, making it a relevant therapeutic target. In another study presented by research groups from the Olomouc node of the NICR, the Cell Painting assay was used to identify compounds that modulate autophagy. Morphological profiling clearly distinguished autophagy inhibitors from activators. Inhibitors showed features consistent with impaired autophagic flux, including Golgi fragmentation, mitochondrial swelling, and ER expansion. Notably, MCOPPB, although known as a NOP receptor agonist, clustered with autophagy inhibitors. This unexpected behaviour of MCOPPB points to a previously unrecognised mechanism and highlights the value of combining morphological profiling with marker-based validation in the search for novel autophagy modulators with potential oncological relevance. (Abstract 6403. DOI: 10.1158/1538-7445.AM2026-6403)
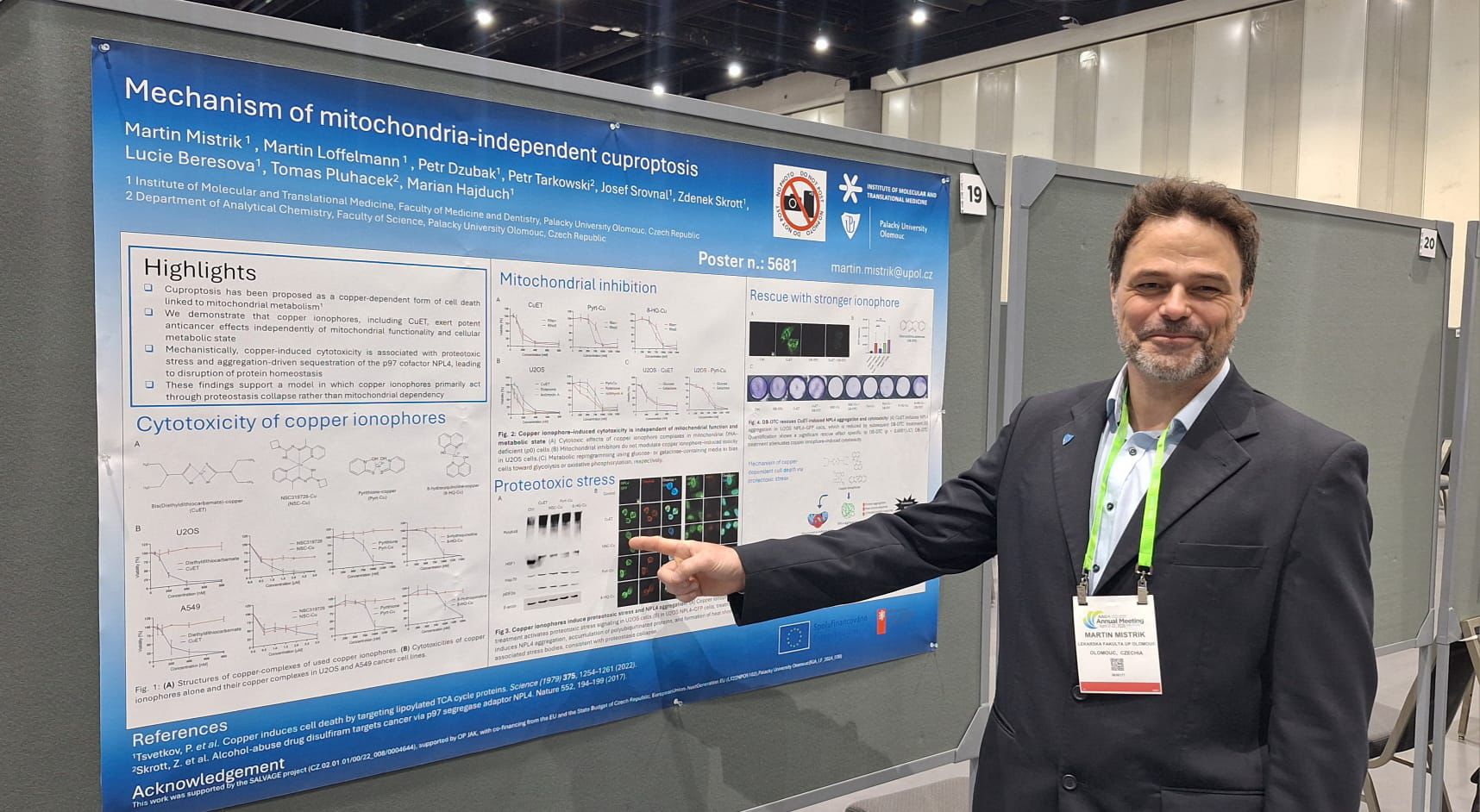
Mechanism of mitochondria-independent cuproptosis.
And thirdly, a presentation of the work of research groups from the Olomouc node of the NICR. Copper is an essential trace element that functions as a cofactor for numerous metabolic and detoxification enzymes. In excess, however, copper exerts marked cytotoxicity in cancer cells and can induce a recently characterized form of regulated cell death termed cuproptosis. Currently, cuproptosis is believed to depend on mitochondrial activity. The authors evaluated several copper ionophores in their complexed forms with copper, including bis(diethyldithiocarbamate) (CuET), pyrithione, NSC319726, and 8-hydroxyquinoline, to assess their ability to induce cuproptosis. Surprisingly, their cytotoxic effects were comparable in oxidative-phosphorylation-dependent cancer cells and in glycolysis-driven counterparts. Consistently, neither chemical inhibition of individual mitochondrial complexes nor the use of mitochondrial DNA-deficient Rho0 cells revealed differential sensitivity. Instead, all tested ionophores induced aggregation and immobilization of the essential p97 cofactor NPL4, mirroring the mechanism previously reported for CuET. Importantly, additional treatment with the non-toxic, more potent divalent copper chelator, dibenzyldithiocarbamate, reversed NPL4 aggregation and ionophore-induced cytotoxicity. Together, these findings refine the mechanistic framework of copper-dependent cell killing by revealing a prominent proteotoxic component of bivalent copper associated with targeting the NPL4. This insight strengthens the rationale for targeting copper-regulated proteostasis pathways as a potential anticancer strategy. (Abstract 5681. DOI: 10.1158/1538-7445.AM2026-5681)
NQO1 as a target to overcome therapy resistance in multiple myeloma
NAD(P)H:quinone oxidoreductase 1 (NQO1) has been linked to poor prognosis and therapy resistance in solid tumours, but its role in multiple myeloma (MM) is less well defined. This study by an international team, which included research groups from the Brno node of the NICR, examined its impact on treatment response and immunotherapy targets in MM using RNA sequencing of CD138+ cells from 24 newly diagnosed and 69 relapsed/refractory patients, alongside MM cell lines.
NQO1 expression was significantly higher in relapsed/refractory MM than in newly diagnosed disease. In MM cell lines, NQO1 overexpression increased resistance to the proteasome inhibitors (PI) bortezomib and carfilzomib, while sensitivity to the immunomodulators (IMiDs) remained unchanged. Pharmacological inhibition of NQO1 with ES936 restored PI sensitivity to wild-type cells levels.
Surface antigen analysis showed no changes in BCMA, SLAMF7 or GPRC5D, but CD38 expression was significantly reduced in NQO1-high cells. Accordingly, these cells were less sensitive to the anti-CD38 antibodies daratumumab and isatuximab, whereas ES936 restored CD38 expression and treatment response. As CD38 mRNA levels were unchanged, the mechanism is likely post-translational.
These findings identify NQO1 as a driver of resistance to both proteasome inhibitors and CD38-directed therapies in MM, and support NQO1 inhibition as a promising strategy in relapsed/refractory disease. (Abstract 3912. DOI: 10.1158/1538-7445.AM2026-3912)
GPRC5D expression on MM cells amplifies the effect of talquetamab
G protein-coupled receptor class C group 5 member D (GPRC5D) is an emerging immunotherapeutic target in multiple myeloma (MM). Talquetamab, a GPRC5D-directed bispecific antibody, has shown approximately 70% efficacy in triple-class refractory MM, although relapse due to genomic loss is frequently observed. To investigate how talquetamab, GPRC5D loss, and T cells interact within the immune microenvironment, an international team of authors – including research groups from the Brno node of the NICR – performed single-cell sequencing in cocultures of healthy T cells with CRISPR-Cas9-generated monoallelic and biallelic GPRC5D knockout MM models, with or without talquetamab.
CITE-seq revealed marked transcriptomic changes in MM cells with both monoallelic and biallelic GPRC5D loss. Activated CD4 and CD8 T-cell populations, defined by canonical activation markers, were detected only in the presence of both talquetamab and GPRC5D-positive MM cells and were absent in cocultures with GPRC5D-null models. Across T-cell subsets, talquetamab in the presence of GPRC5D-positive MM cells induced strong upregulation of interferon-stimulated genes, including GBP2/4/5, IFI44, IFIT2, and ISG15. Notably, talquetamab alone also increased expression of interferon-signalling genes such as STAT1 and IRF1, even without GPRC5D-positive MM cells.
Additional genes associated with T-cell activation, including SOCS3, CD69, JUNB, BATF, CISH, and PRDM1, were upregulated only when both talquetamab and GPRC5D were present, indicating a distinct activation state. Gene set enrichment analysis (GSEA) showed enrichment of IFN-γ and IFN-α signalling across multiple T-cell subsets, whereas IL2-STAT5 signalling and inflammatory response pathways were enriched only in the presence of both talquetamab and GPRC5D-positive MM cells. In the absence of talquetamab, T-cell transcriptomes were unaffected by GPRC5D status alone.
These findings indicate that talquetamab induces interferon-related transcriptional programmes in T cells even without GPRC5D, but this response is amplified when MM cells express GPRC5D. Talquetamab also reshapes T-cell activation states and the distribution of T-cell subpopulations according to the GPRC5D status of MM cells. (Abstract 5550. DOI: 10.1158/1538-7445.AM2026-5550)
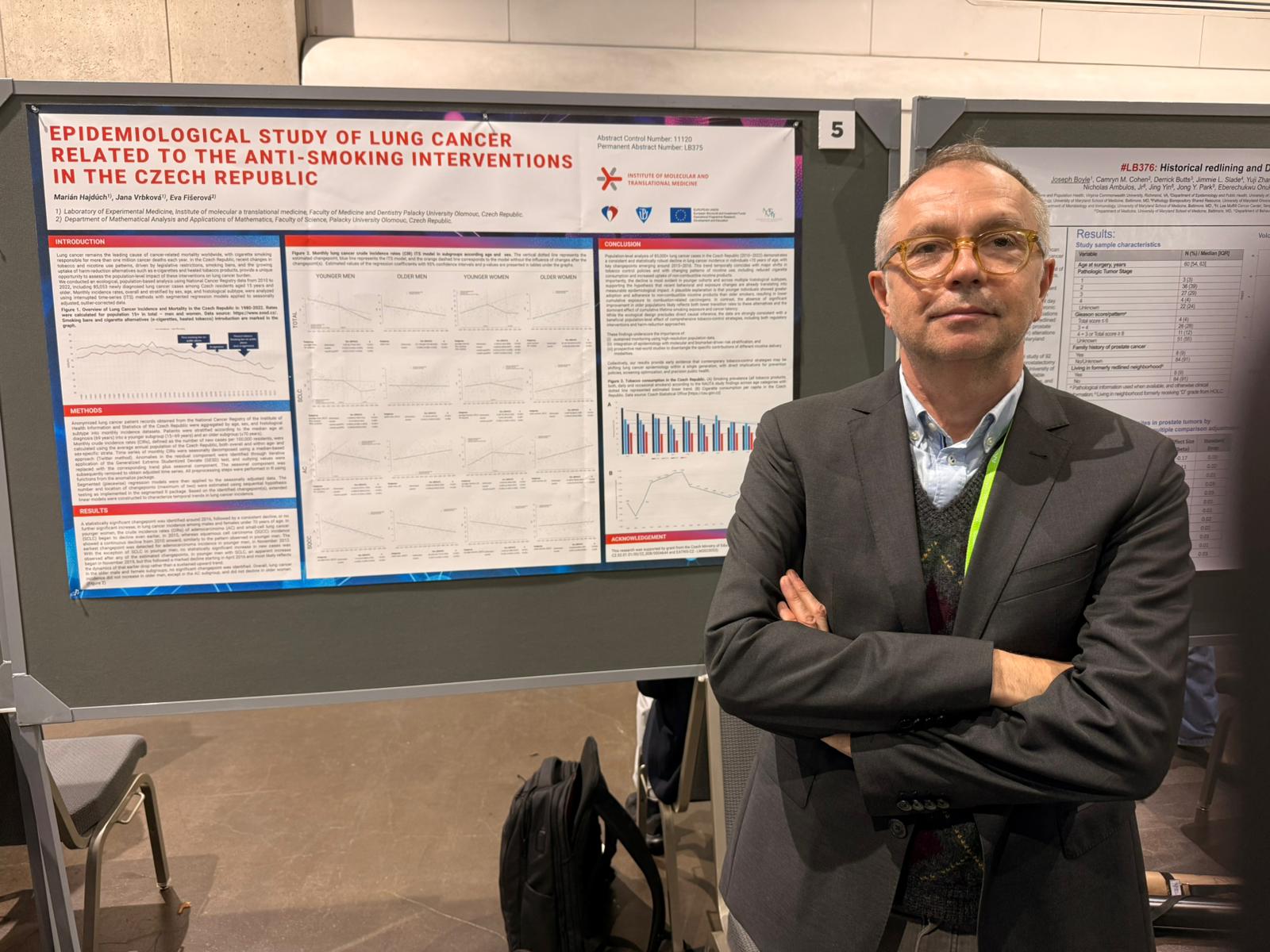
Abstract LB375: Ecological time-series study of lung cancer incidence in relation to tobacco control and harm reduction in the Czech Republic
The latest of the selected posters is once again from the Olomouc node of the NICR and was presented in the Late Breaking Abstracts section. Lung cancer remains the leading cause of cancer mortality and is largely attributable to tobacco smoking. In the Czech Republic, tobacco-control policies, smoking behaviour, and the use of smoke-free nicotine alternatives have changed over the past decade. This ecological time-series study analysed National Cancer Registry data from January 2010 to December 2022, including 85,065 incident lung cancer cases in individuals aged 15 years or older, to assess overall and sex-, age-, and histology-specific incidence trends, including small-cell lung cancer (SCLC), non-small cell lung cancer (NSCLC), adenocarcinoma (AC), and squamous cell carcinoma (SQCC). Changepoints were selected using the Bayesian Information Criterion, and post-changepoint trends were compared with counterfactual projections based on pre-changepoint slopes.
Overall incidence remained broadly stable, but this masked divergent sex-specific patterns, with a steady decline in men and an increase in women. SQCC declined markedly, with a breakpoint around March 2013, and SCLC decreased mainly in men, whereas women, particularly older women, showed stable or rising SCLC incidence. The strongest age-by-sex divergence was observed for AC: incidence continued to rise in older women, while in younger women it peaked around July 2015 and subsequently declined or stabilised relative to counterfactual growth.
The decline in younger women may plausibly relate to greater uptake and sustained use of smoke-free nicotine alternatives, resulting in lower exposure to combustion-related carcinogens, although this interpretation is necessarily hypothesis-generating given the ecological design. Because AC may arise through both smoking-dependent and smoking-independent pathways, and causal inference is limited at the population level, longer follow-up and linkage with individual exposure data are needed. (Abstract LB375. DOI: 10.1158/1538-7445.AM2026-LB375)